BIBI IV - One click mode How-To
Query
SSU-rDNA 16S Fasta Sequence
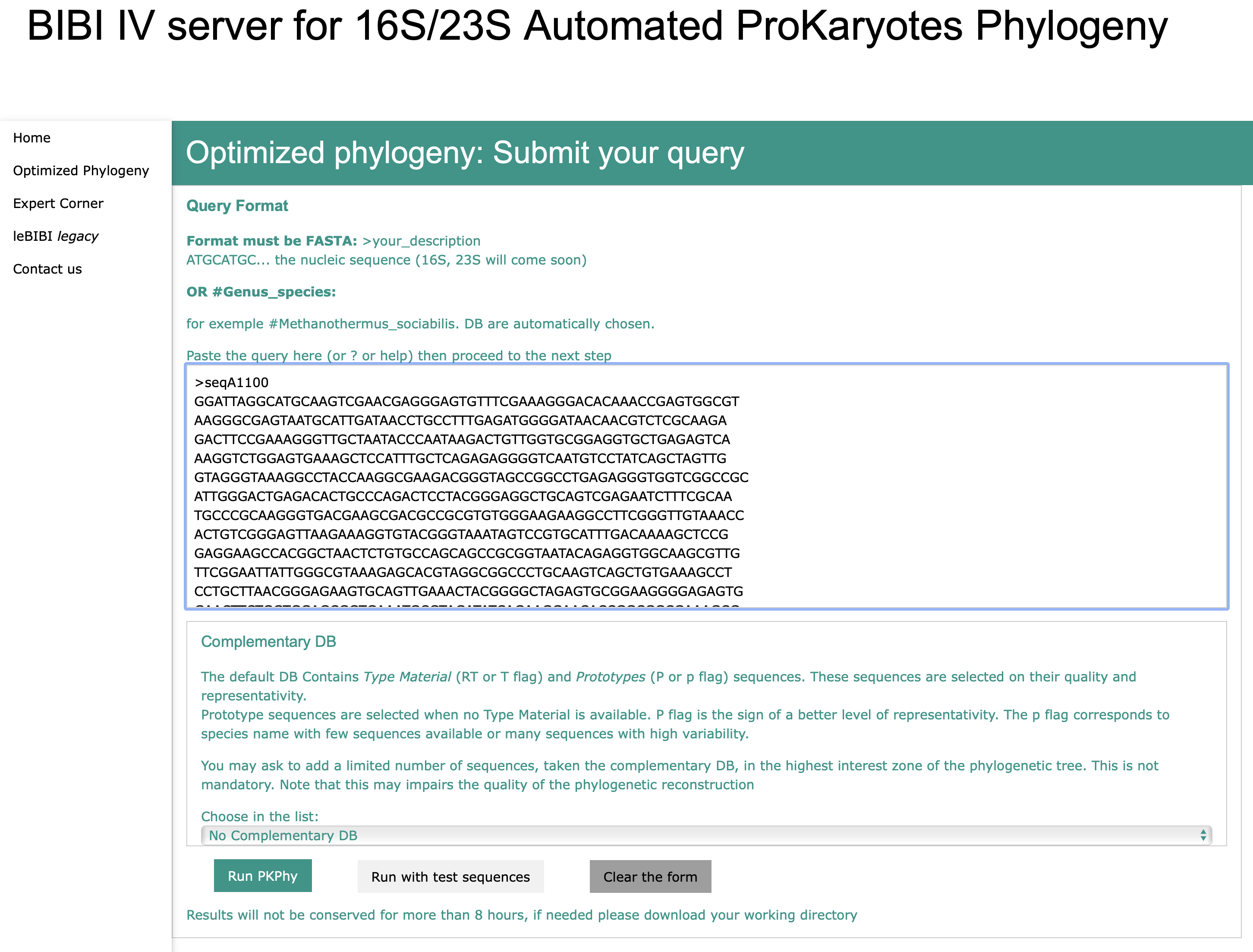
Species Name
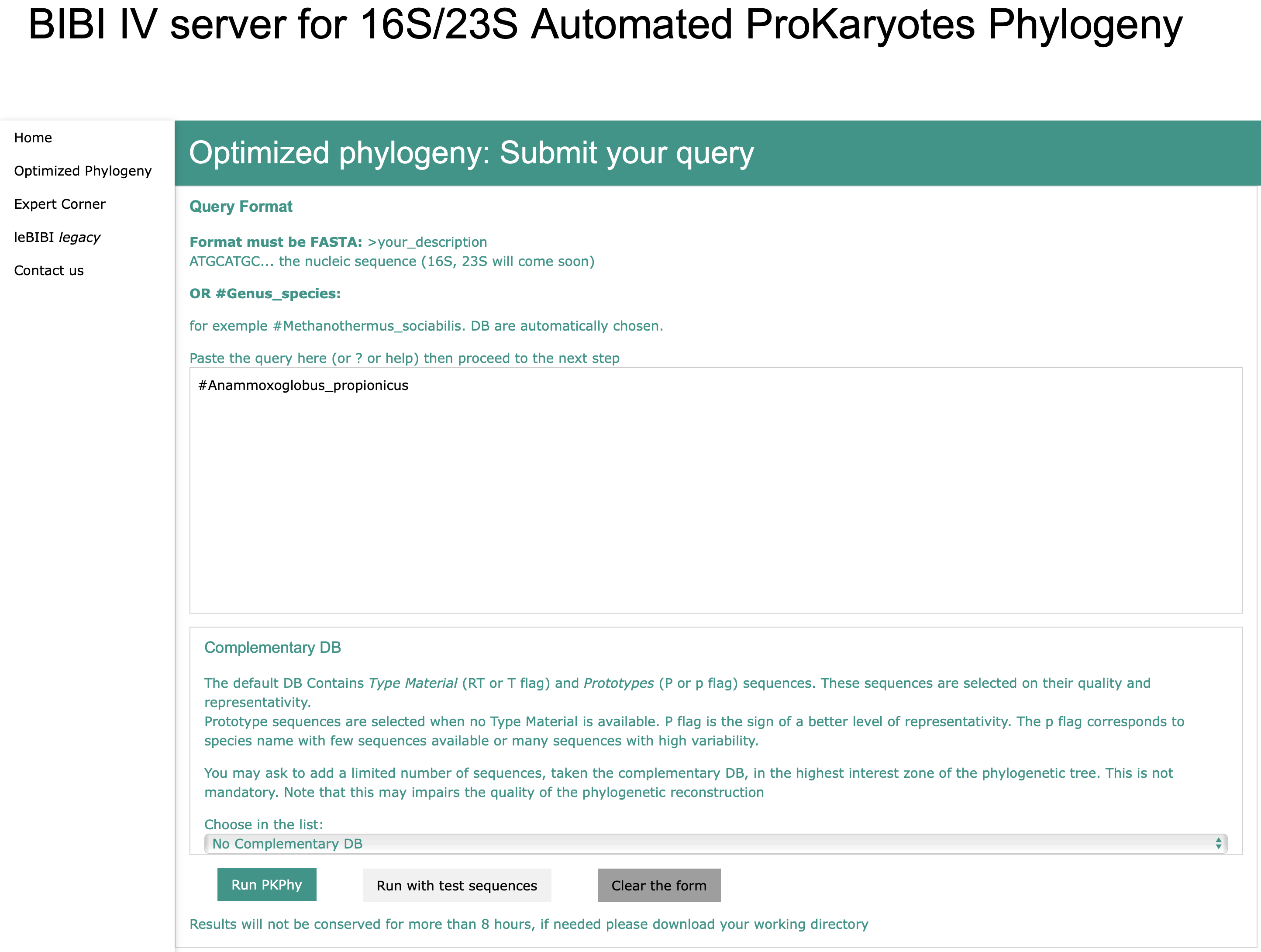
Selection of the complementary Database
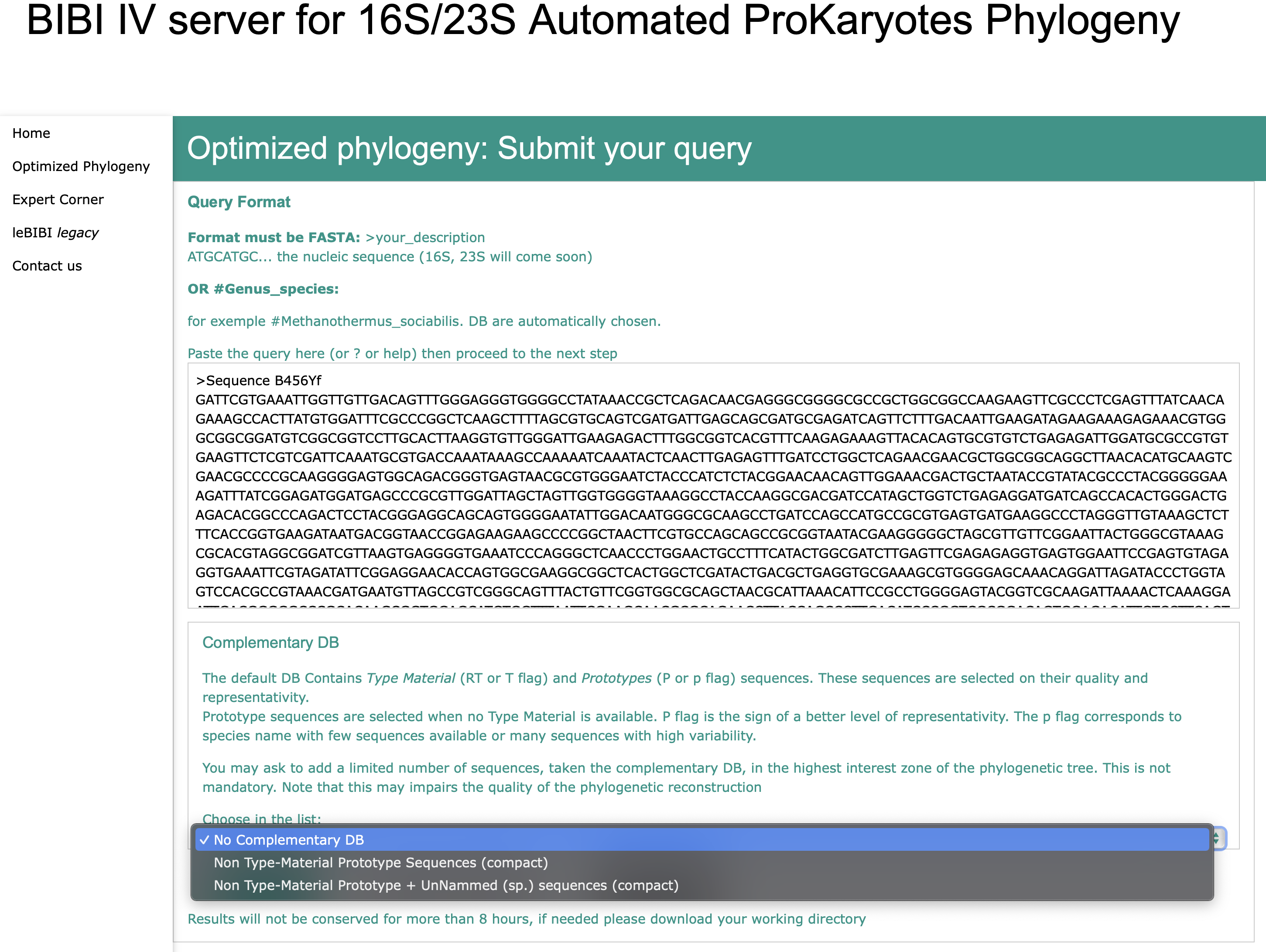

Send to BIBI IV
Run (or test)
Running

Intermediate result (important if multiple queries)
Intermediate result Wait for Complementary DB
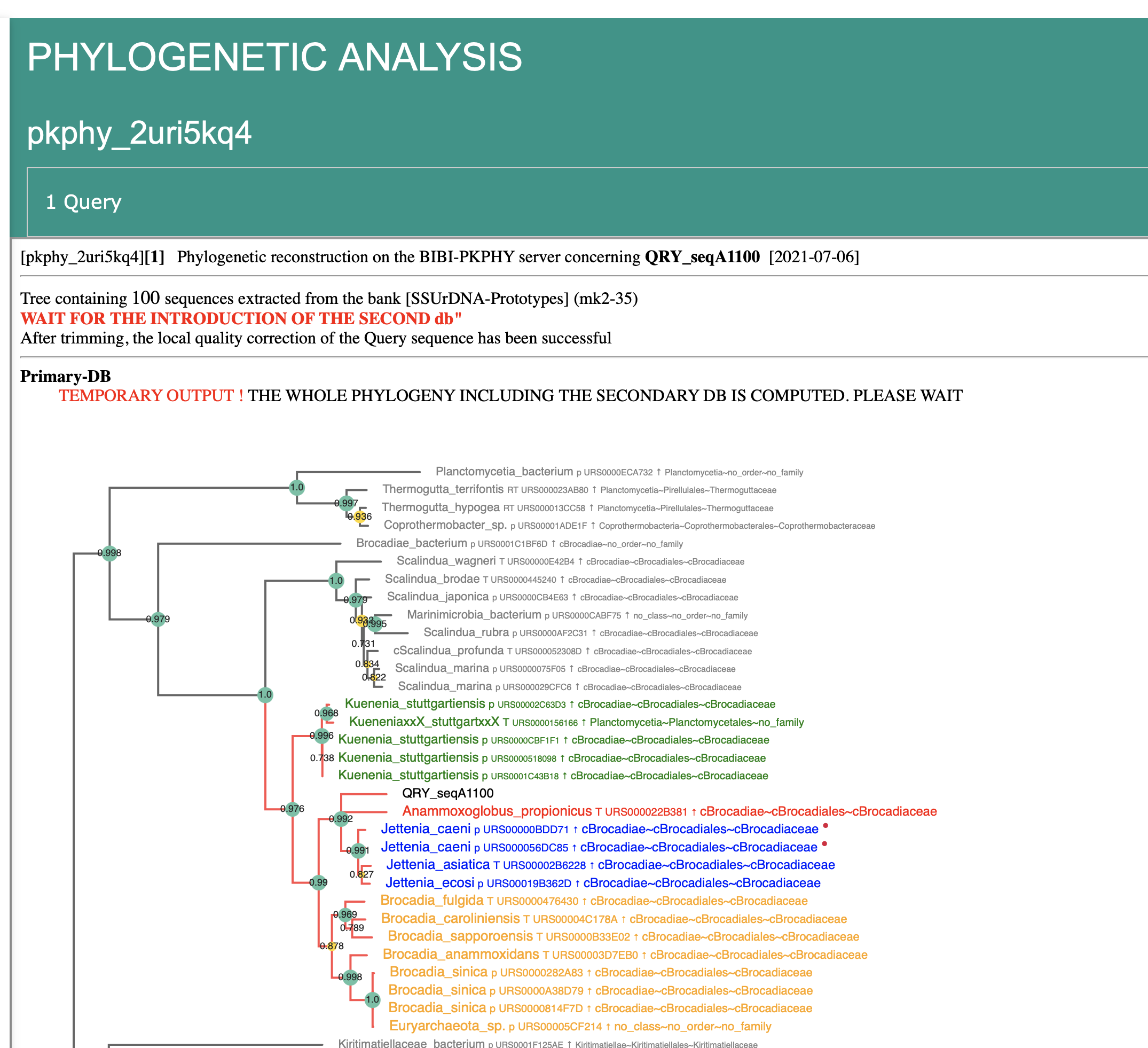

Results
Final menu - multiple queries

Final menu - one query
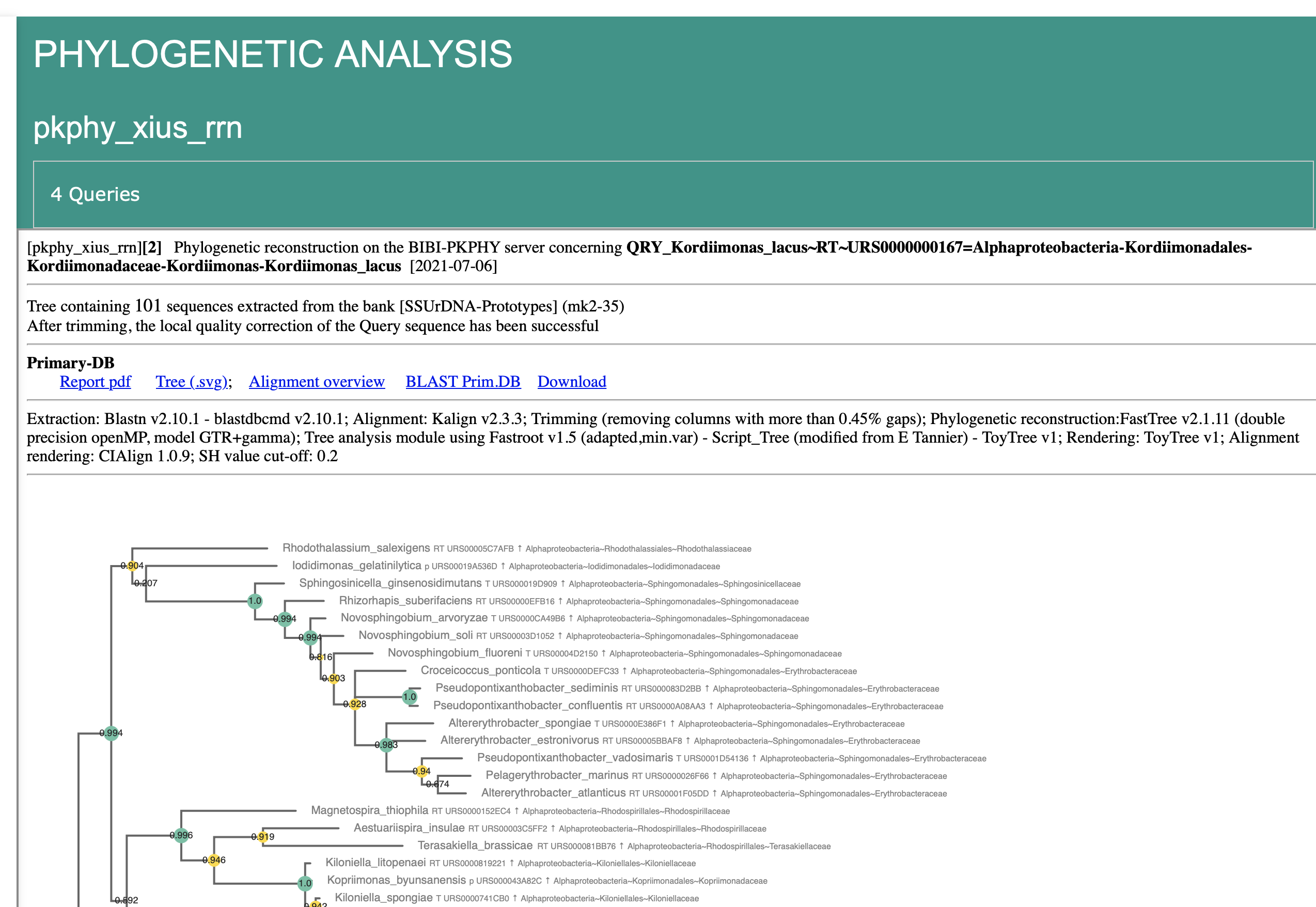
Individual Query Result
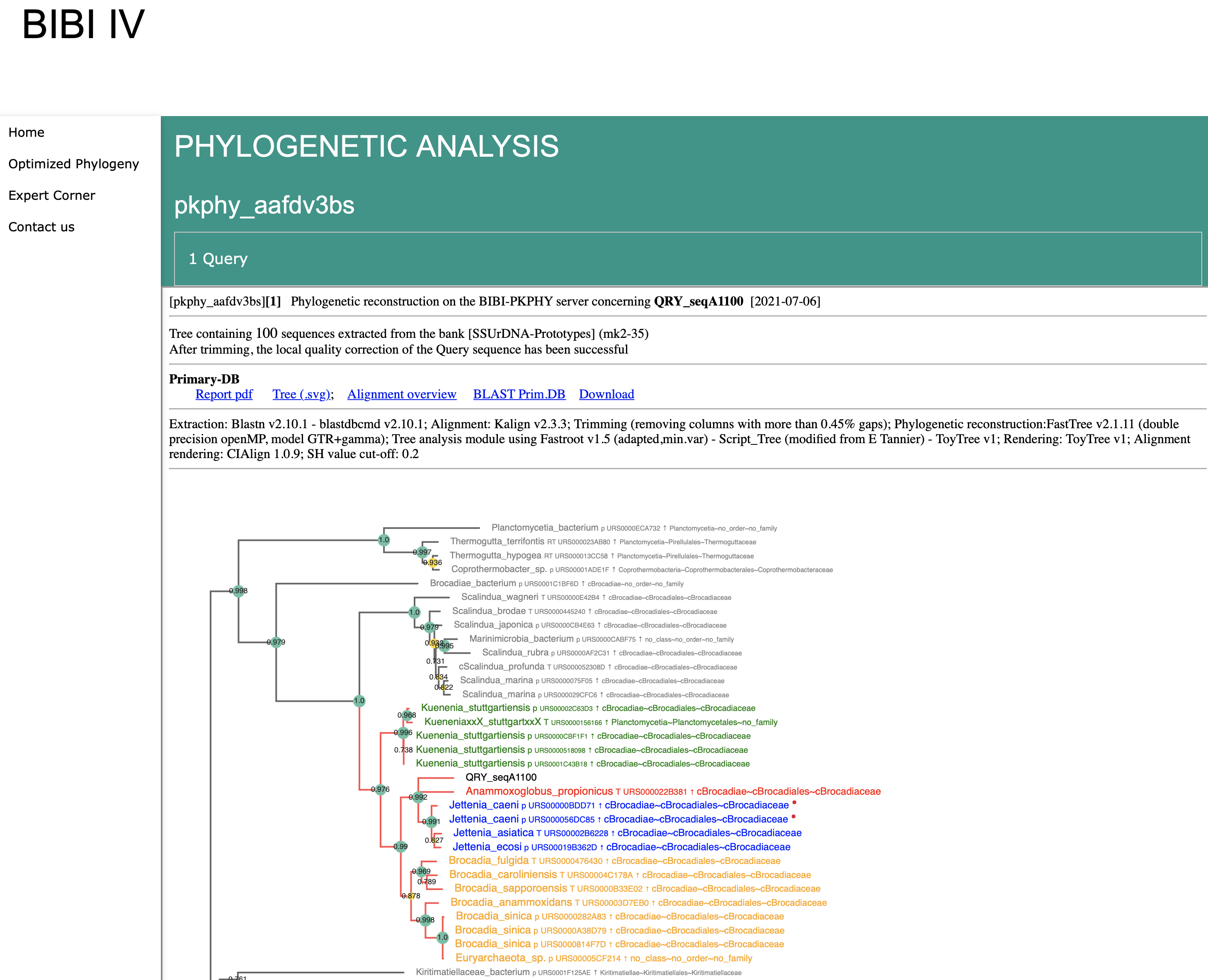

Summary of the methods
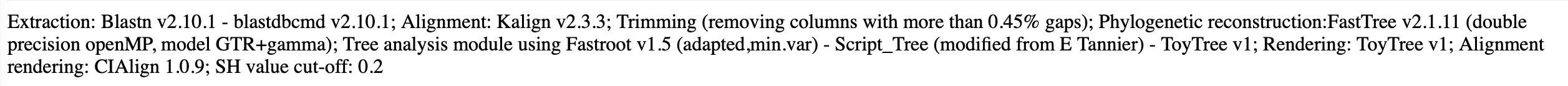
Individual Query Tree Features
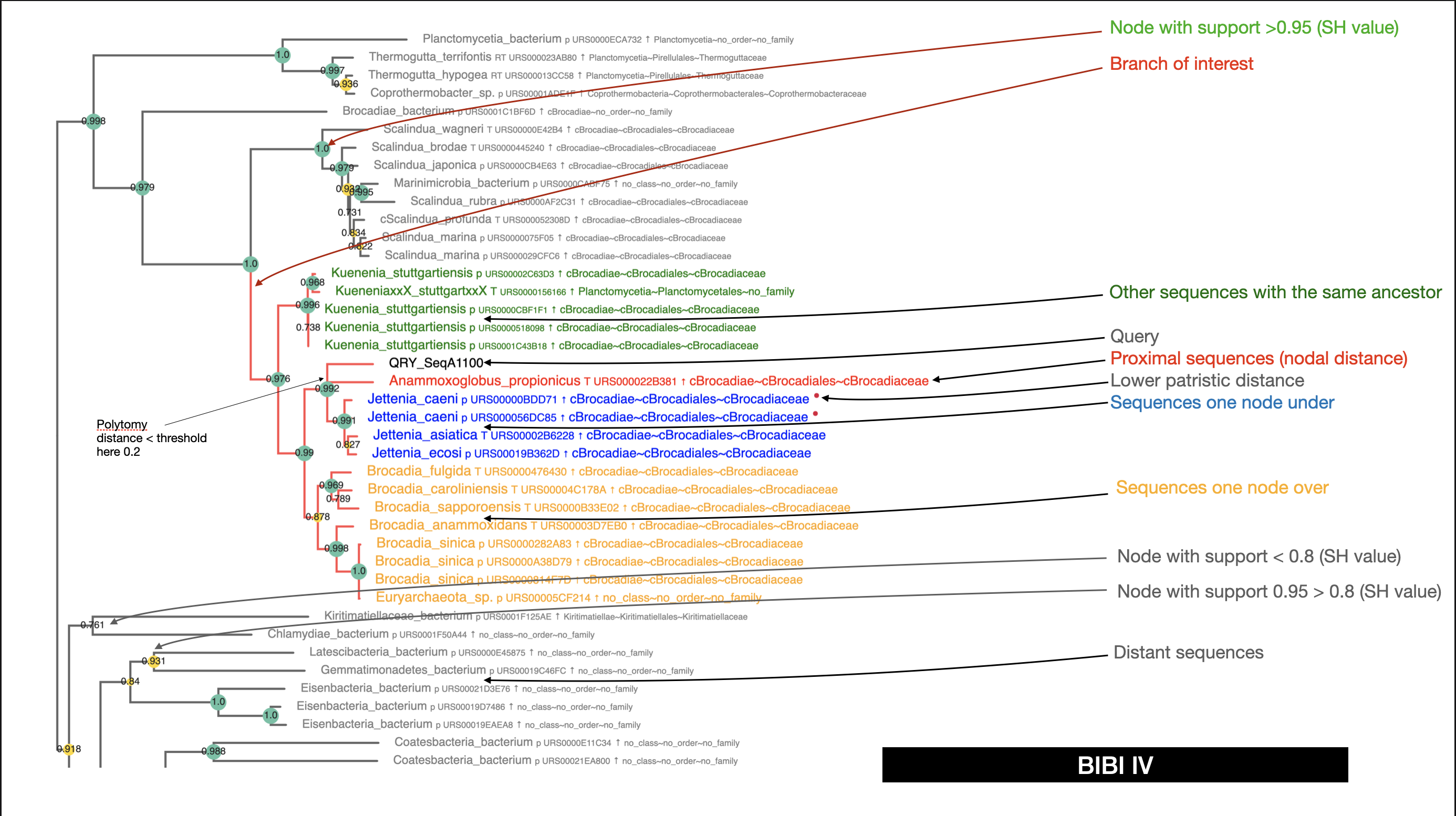
Branch labels and links
This is an example of branch label:

Here is an exemple with the same links (active) as found in the tree:
Methanothermus_fervidus RT URS000014A460 ↑ Archaea~Euryarchaeota~Methanobacteria
You may discover the three links to LPSN, RNAcentral and DDBJ taxonomy server.
Clicking on ↑ will launch a new phylogeny by taking the species name as query, in this case the number of leaves is 150 to extend the exploration (primary DB only).
Note that the hierarchy is limited to Class-Order-Family as the Genus is given by the first part of the species name and is not repeated to prevent too long descriptions. Note that "candidatus" is replaced by a "c" prefix to the name, ex: cZinderia_insecticola. This is not following the Nomenclature code but this helps the computer treatment and these "candidatus" are de facto recognized by the community, if not by the official nomenclature. In this exemple, TR is related to the Sequences classification.
Nodes
Nodes having a SHvalue less or egal to 0.2 are replaced by a multifurcation. Nodes are yellow when the SH value is between 0.8 and 0.95 and green when SH value is over 0.95. The diameter of the yellow node is also proportional to the SH value within [0.8:0.95[
So, we can be confident in the green nodes ( Colors signification) and interpretate the relations accordingly. SH values > 0.95 are currently admitted as significative support value, 0.9 being more challenging.
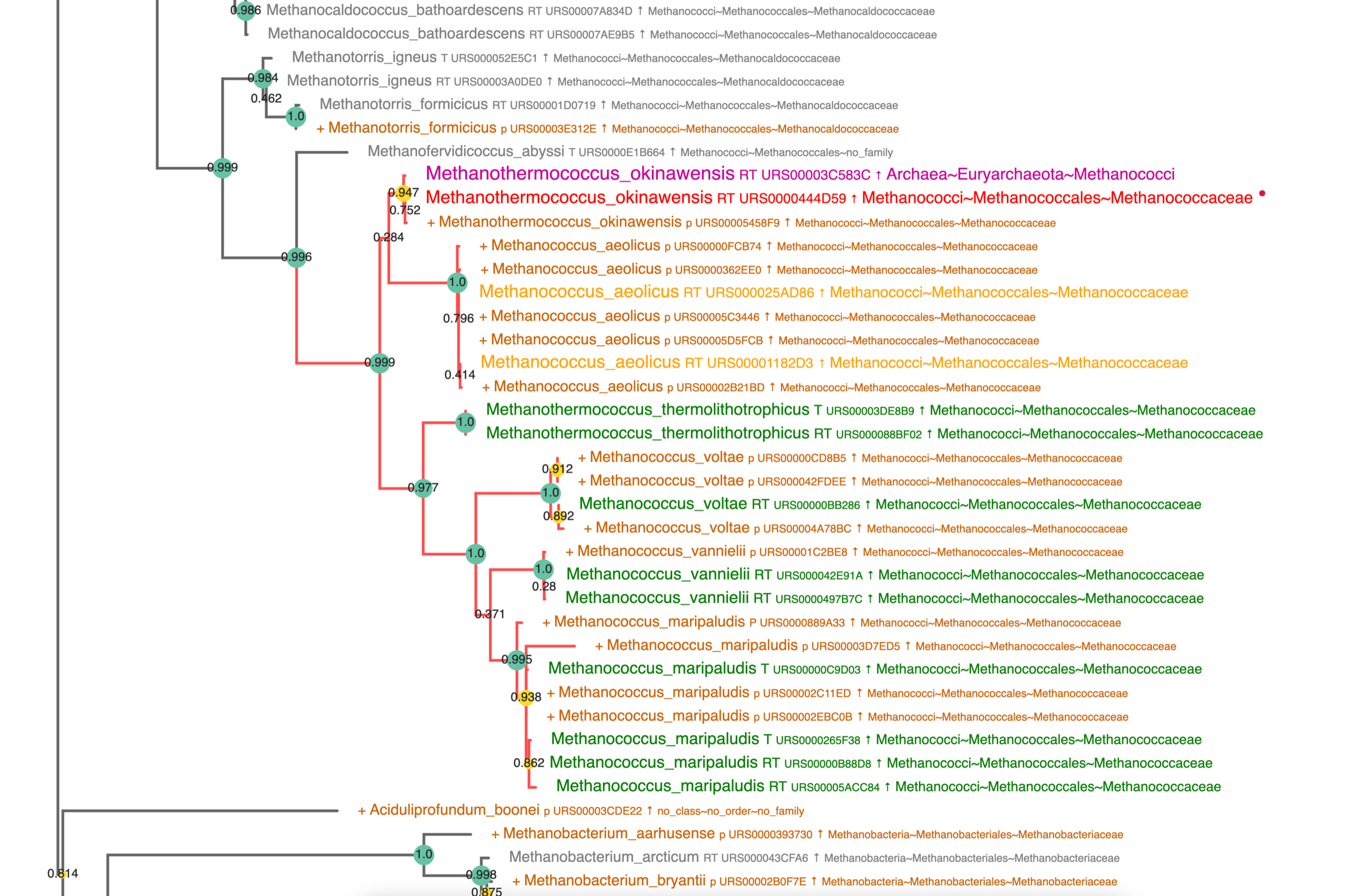
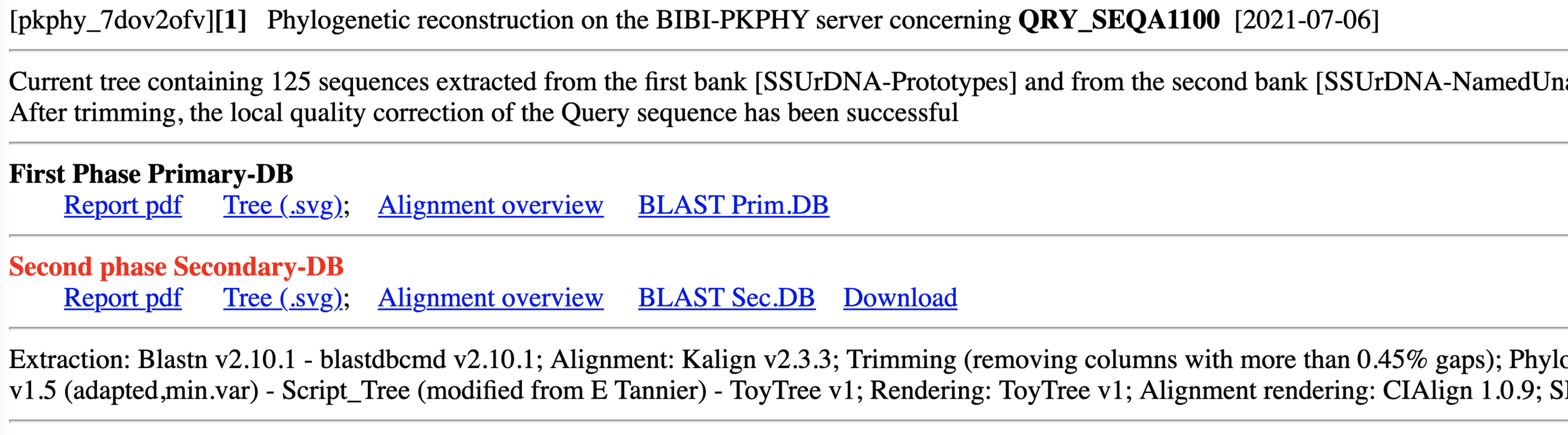
Individual Query Result Two Banks
Note the "+" in front of the sequence identification for the complementary bank !

Here a centered view of the tree: the added sequences enhance the diversity.
Files and Downloading

Troubleshooting
If the process stops
Please try to find your pkphy identifier (pkphyXXXXX like pkphy__dbe9rc7) in The Users Directory.
Usually it is clean :
If the process stops the directory is full of technical files and logs

Please copy the logsERR.txt file and send it along with your query to: us